
Research teams led by Greg Moorhead and Ken Ng investigated an enzyme named RLPH2.
Riley Brandt, University of Calgary
April 23, 2018

Research teams led by Greg Moorhead and Ken Ng investigated an enzyme named RLPH2.
Riley Brandt, University of Calgary
Biologists in the Faculty of Science have discovered a new and unique, enzyme-controlled cell-signalling mechanism in plants, which enhances understanding of how cells in all living organisms communicate with each other.
Research teams led by Greg Moorhead and Ken Ng at the University of Calgary investigated an enzyme named RLPH2, first identified by Moorhead’s research group.
RLPH2 is a protein phosphatase, a group of enzymes that catalyze a vital chemical reaction, called dephosphorylation, in cells.
In a multidisciplinary collaboration between biochemistry and structural biology, the teams discovered that RLPH2 carries out dephosphorylation in an unusual and previously unknown way.
Protein phosphorylation regulates most aspects of cell life. It is the "conductor" that orchestrates the sending and interpreting of messages within and between cells, by chemically altering thousands of proteins within cells.
This is the dominant way that cells in all living organisms, including humans, communicate in controlling many activities inside cells — and thereby the physiological outcomes in plants, animals and other organisms.
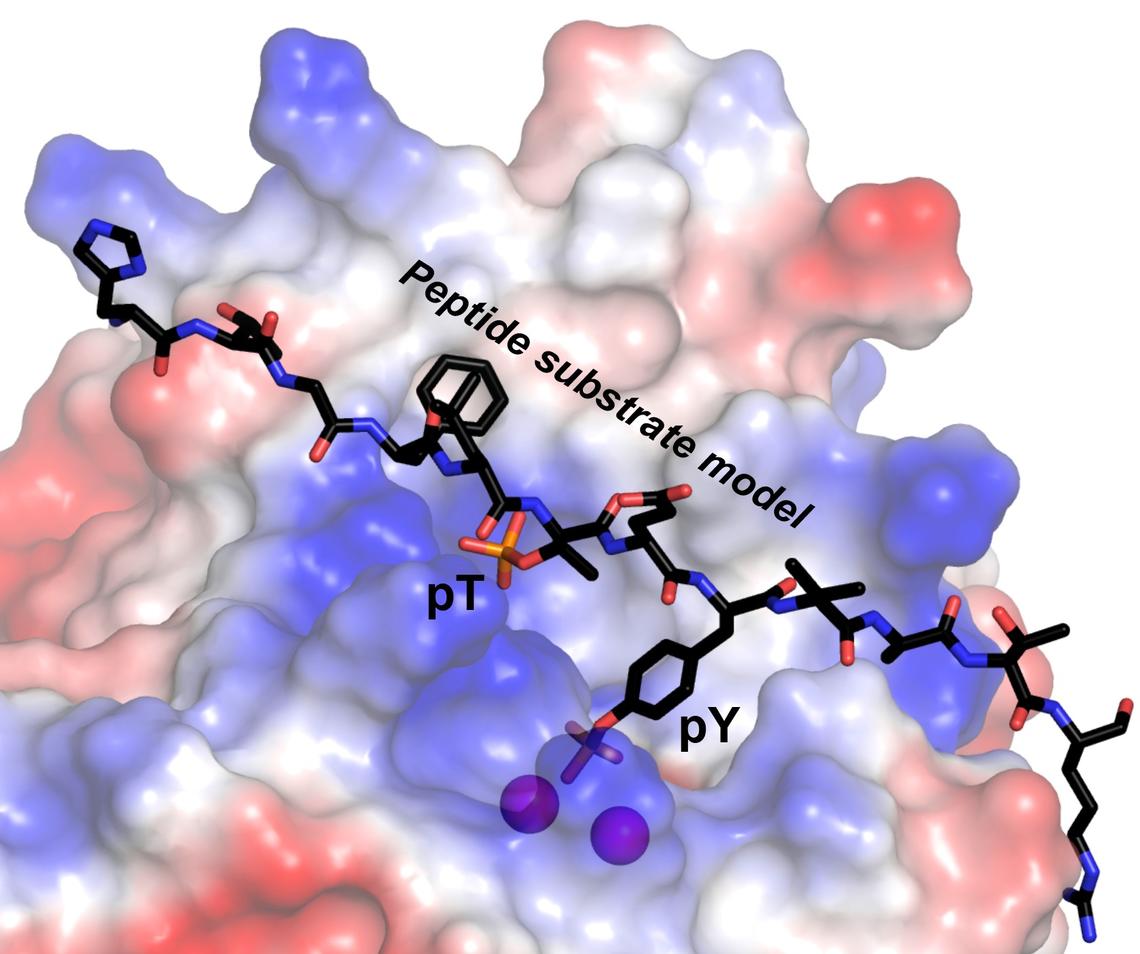
The research teams found that the RLPH2 enzyme has an unusually deep active site structure, with two different “phospho-binding” sites where the enzyme chemically binds with a “substrate” (a molecule upon which the enzyme acts). RLPH2’s two sites include a novel secondary phospho-binding "pocket."
“This enables RLPH2 to carry out very specific dephosphorylation via a dual mechanism, recruiting substrates through the secondary pocket, then ultimately dephosphorylating a nearby amino acid called phosphotyrosine,” says Moorhead, professor of biochemistry in the Department of Biological Sciences.
“The uniqueness of this enzyme is the tyrosine dephosphorylation and the secondary binding pocket, which is completely unique for this group of enzymes,” he says.
“The deeper active site structure allows the enzyme to uniquely recognize the phosphotyrosine, which ‘reaches’ into the active site,” says Ng, professor of structural biology in the Department of Biological Sciences.
The team’s paper, “Structural Basis for the Preference of the Arabidopsis thaliana Phosphatase RLPH2 for Tyrosine-phosphorylated Substrates,” is published in Science Signaling, a journal in the top-ranked Science series.
Cell signalling involved in major human diseases
Cell signalling activated by protein phosphorylation is a "hot" research topic. Abnormal protein phosphorylation is a cause or consequence of major diseases, such as cancer, diabetes and rheumatoid arthritis.
Also, defects in genes that encode the enzymes (protein kinases and phosphatases) involved in phosphorylation underlie several inherited and acquired disorders, such as leukemias, lymphomas and immune diseases.
Almost every large pharmaceutical company and many biotechnology firms are targeting protein kinases and protein phosphatases in developing therapies for diseases, particularly cancers. Such applications wouldn’t be possible without first thoroughly understanding signal transduction and phosphorylation.
“We want to understand all of the protein kinases and phosphatases inside a cell and how they work in each signalling pathway — that’s the ultimate goal,” Moorhead says.
Anne-Marie Labandera led the work of producing protein and growing microscopic-sized crystals.
Revealing the RLPH2 enzyme’s structure
The researchers used bioinformatics, or genome sequencing analysis, to first pinpoint the RLPH2 enzyme, which exists in plants but not in animals.
Moorhead’s team performed the gene cloning and the protein expression (producing the RLPH2 enzyme) and the enzyme assays.
Much of this biochemistry work was done by Anne-Marie Labandera, then a PhD student with Moorhead, and the lead author on the new paper. She’s now in the U.K. doing post-doctoral work at the University of Birmingham.
“Employing a multidisciplinary approach on my PhD was key to characterize this unique plant protein and to understand its atypical behaviour,” Labandera says.
The researchers expected RLPH2 would — like other enzymes in the same group — biochemically target and dephosphorylate phosphoserine and phosphothreonine. These are the two amino acid residues most commonly involved in protein phosphorylation.
To their surprise, RLPH2 preferred to target phosphotyrosine. Tyrosine is involved in only about two per cent of protein phosphorylation in living organisms.
“Based on genomics, there’s only one known tyrosine phosphatase in plants,” Moorhead says. “So I think this RLPH2 enzyme is really important to add to that group, and in part explains this lack of tyrosine phosphatases in plants because this other enzyme has been adapted to play that role.”
Ng and his team used the protein produced by Moorhead’s laboratory to grow microscopic-sized crystals. They then shone X-rays on the crystals, from which they mathematically calculated RLPH2’s 3-D structure — down to individual molecules.
“The structure tells us how this enzyme is able to recognize some little piece of a protein and do its activity on just that,” Ng says.
Collaboration key to the discovery
The unique specificity of RLPH 2 suggests it controls the activity of one or more of a group of important enzymes in the cell called mitogen activated protein kinases, or MAPKs, Moorhead says. “The MAPK enzymes receive information from a variety of sources, perpetuating the signal and allowing the cell to respond appropriately to the information being sent.”
Without the collaboration between the biochemistry and structural biology teams, “this discovery wouldn’t have happened,” Moorhead says.
Labandera agrees, saying: “Thanks to this collaboration with Prof. Ng, we figured out the necessary features that RLPH2’s targets need to possess, for ultimately understanding its physiological relevance.”
Adds Ng: “The strength of the Faculty of Science is we’re all very interested in these fundamental research questions. And from this fundamental research, you can’t tell — 10, 20 or 30 years later — where the big applications will come.”
The teams’ research was supported by the Natural Sciences and Engineering Research Council of Canada.